Every year, millions of people around the world fall victim to sepsis—a life-threatening reaction to infection that can overwhelm the body and shut down its organs. Doctors often describe treating sepsis as a “race against time.” And with good reason: for every hour treatment is delayed, survival rates drop dramatically. According to clinicians, patients in septic shock lose nearly 8 percent of their chance of survival with each passing hour.
Yet today’s diagnostic tools move painfully slowly. The standard method used in hospitals—blood culture testing—requires waiting for bacteria to grow in a lab dish. That process usually takes a day or more, sometimes up to four days before doctors can be sure which bacteria are responsible. In those critical hours, the infection can spread rapidly, leaving doctors with little choice but to administer broad-spectrum antibiotics. While these drugs can save lives, they also bring serious risks, from damaging gut bacteria to fueling the global crisis of antibiotic resistance.
But what if doctors could confirm a bacterial infection within just a couple of hours, and identify the pathogen in less than half a day? That is the promise of a groundbreaking diagnostic method developed by researchers at KTH Royal Institute of Technology and Uppsala University, published recently in npj Digital Medicine.
A Smarter Way to Catch Sepsis Early
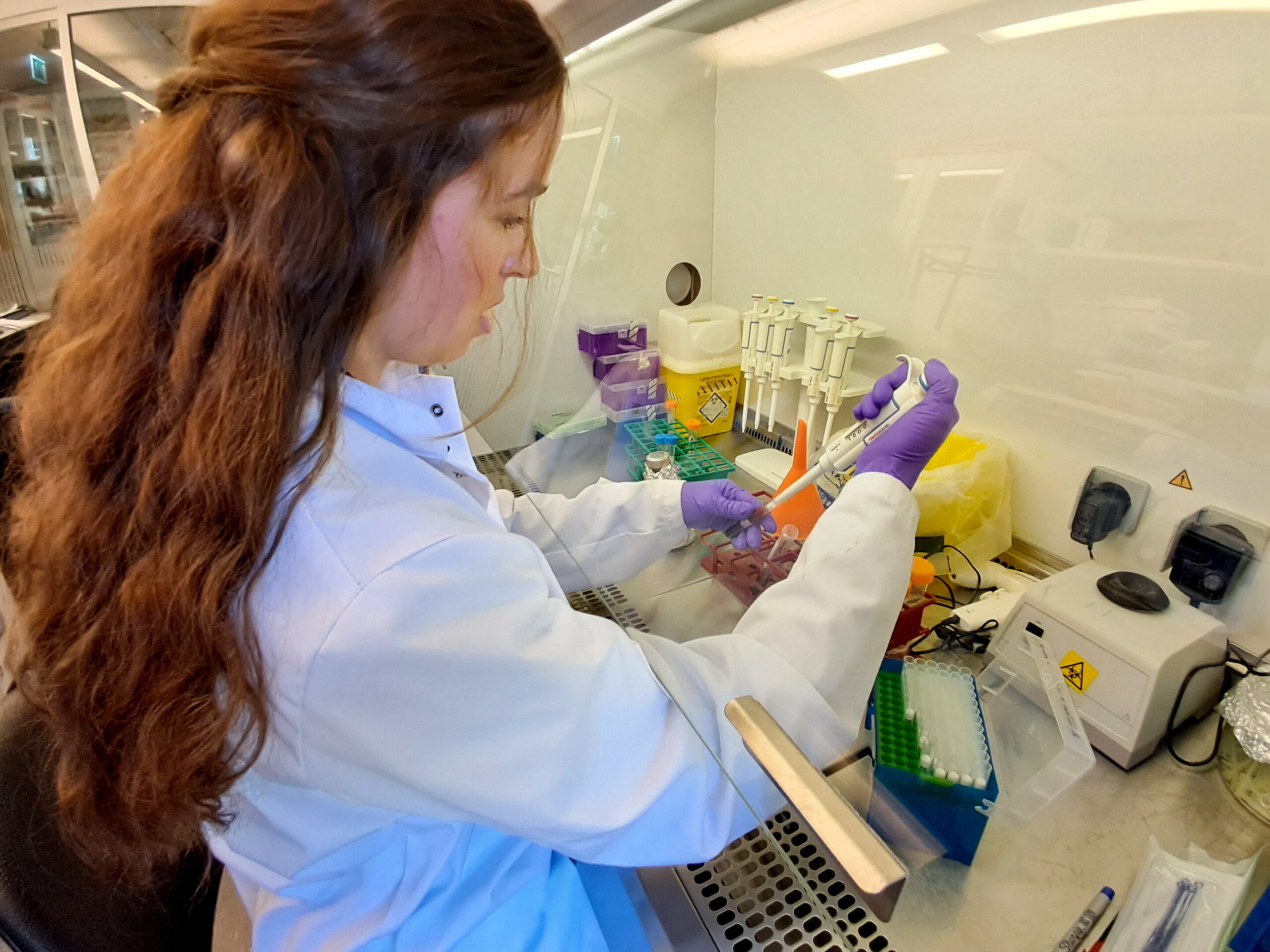
The new process, spearheaded by doctoral researchers Henar Marino Miguelez and Mohammad Osaid, replaces days of anxious waiting with a window of just two to six hours. Instead of relying on bacterial growth, it physically isolates and detects bacteria in blood samples using advanced tools: smart centrifugation, microfluidic chips, automated microscopy, and artificial intelligence.
Here’s how it works. A patient’s blood sample is spun in a centrifuge along with a special agent. This causes the blood cells to sink downward while bacteria float into a clear liquid layer above. From there, the bacteria-rich fluid is funneled into a microchip etched with tiny channels. Hidden in these channels are microscopic traps designed to catch bacteria. Once captured, the bacteria begin to grow, and time-lapse images taken under an automated microscope reveal their presence.
Artificial intelligence then analyzes the images with remarkable sensitivity, capable of detecting even tiny bacterial populations—just 7 to 32 colony-forming units per milliliter of blood in laboratory tests. The system has already proven its ability to detect pathogens like Escherichia coli, Klebsiella pneumoniae, and Enterococcus faecalis, all common culprits in dangerous bloodstream infections.
Why Every Hour Matters
For clinicians, this speed is revolutionary. “Diagnosing sepsis is a race against time,” says Marino Miguelez. “With every hour of delayed treatment of patients in septic shock, survival rates drop by 8 percent.”
By identifying bacteria in only a few hours, hospitals could administer the right antibiotic almost immediately instead of relying on broad-spectrum drugs. This not only improves patient outcomes but also curbs the dangerous overuse of antibiotics. As Professor Wouter van der Wijngaart, who leads research in microfluidic and biomedical systems at KTH, explains: “It takes a hospital two to four days before they are sure which antibiotic to treat a bloodstream infection with. We’re trying to do this in four to six hours.”
The potential impact is enormous. Earlier diagnosis could prevent countless deaths, shorten hospital stays, and reduce the long-term complications that sepsis survivors often endure.
Challenges and Next Steps
Of course, no breakthrough is without its hurdles. One significant challenge is Staphylococcus aureus, a bacteria that often causes sepsis but is notoriously difficult to isolate because it hides within blood clots. The team acknowledges this limitation and is already working on refinements to capture and detect it more effectively.
Another step is moving from laboratory testing into clinical environments. The technique has proven successful in spiked blood samples, but widespread use in hospitals will require rigorous clinical trials, regulatory approval, and eventually, integration into healthcare systems.
Still, the promise is clear: a diagnostic tool that could transform how doctors fight sepsis, turning hours of uncertainty into hours of action.
A Glimpse into the Future of Medicine
This breakthrough is more than just a faster test—it represents a larger shift in how technology is reshaping medicine. By combining clever physical techniques like centrifugation with cutting-edge artificial intelligence, researchers are building tools that see what the human eye cannot. These tools promise not only speed, but precision—treating the right patient with the right drug at the right time.
The collaboration between KTH and Uppsala University highlights another truth: science thrives when disciplines meet. Microfluidics, biology, computing, and medicine all came together in this work, proving once again that the future of healthcare depends on innovation across boundaries.
As the fight against antibiotic resistance intensifies and the burden of sepsis continues to weigh on hospitals worldwide, this new diagnostic approach offers hope. Every life saved is not just a victory for science, but for humanity.
And perhaps one day soon, when a patient arrives at an emergency ward with the early signs of sepsis, doctors will not have to wait in anxious uncertainty. They will have answers within hours, and with them, the power to act decisively.
More information: M. Henar Marino Miguélez et al, Culture-free detection of bacteria from blood for rapid sepsis diagnosis, npj Digital Medicine (2025). DOI: 10.1038/s41746-025-01948-w